Interview with Professor Russell Vance
By Matthew Colbert and Akash Kulgod
Dr. Russell Vance is a Professor of Molecular and Cell Biology at UC Berkeley. Professor Vance’s research interests focus largely on the initial encounters between pathogens and the immune system in the context of inflammasomes, Legionnaires’ disease, and tuberculosis. In this interview we examine survival mechanisms of Legionella pneumophila, the pathogen that causes Legionnaires’ disease and the influence of tuberculosis on our cells. We also discuss how the study of one disease can influence research on another.
BSJ: A lot of your work focuses on Legionnaires’ disease. What drove you to focus your research on Legionnaires’ disease and, more recently, tuberculosis?
RV: Tuberculosis is a major global problem—millions of people every year are infected and we currently lack effective vaccines to prevent this. Moreover, many of the antibiotics that we use to treat tuberculosis are increasingly rendered ineffective as the pathogen acquires resistance to them. We’re in desperate need of new therapeutic approaches to this global problem. There’s a great opportunity for scientists like myself to try and understand more about how this pathogen causes disease and develop new therapeutic approaches. I have been working on Legionnaires’ disease for a long time. Legionella is not a major global pathogen, but it’s still a very useful model pathogen that has taught us a lot about how the immune system defends against bacteria. My hope is that some things that we learn about how the body effectively deals with Legionella can be helpful in developing approaches to dealing with tuberculosis. For the most part, people are very resistant to Legionnaires’ disease but are more susceptible to tuberculosis. It is important to figure out what makes people resistant to Legionella and why those defenses don’t work in the case of tuberculosis.
BSJ: Legionnaires’ disease is caused by Legionella, an auxotrophic bacterium that does not make amino acids and cannot reproduce on its own. Without self-replication, how does Legionella replicate?
RV: Legionella can make some amino acids, it just can’t make all of them—so it’s auxotrophic for many amino acids. What that means is that Legionella is almost always replicating within some kind of host cell from which it can acquire these necessary amino acids. We know about Legionella infections because they’re the cause of Legionnaires’ disease—that’s when the bacteria replicate in our lungs. Legionnaires’ disease was only discovered in 1976. But for millions of years before that, Legionella was replicating out in the environment in other host cells like single-celled amoebae, and probably never wanted to have anything to do with humans. We often regard Legionella as an accidental pathogen in humans, as humans generally acquire it by breathing in large doses that have been aerosolized by cooling towers on the top of buildings. Only because of the invention of such towers did humans ever get the disease.
BSJ: One of your papers mentions that the protein mTORC1 regulates protein production in the cell. How is it impacted by Legionnaires’ disease?
RV: mTORC1 is a master regulator of amino acid metabolism in humans. What a student in my lab, Justin De Leon, discovered is that Legionella encodes bacterial factors that manipulate the activity of mTORC1. Our hypothesis is that Legionella does that to trick the host into producing amino acids that Legionella can consume for its own benefit.
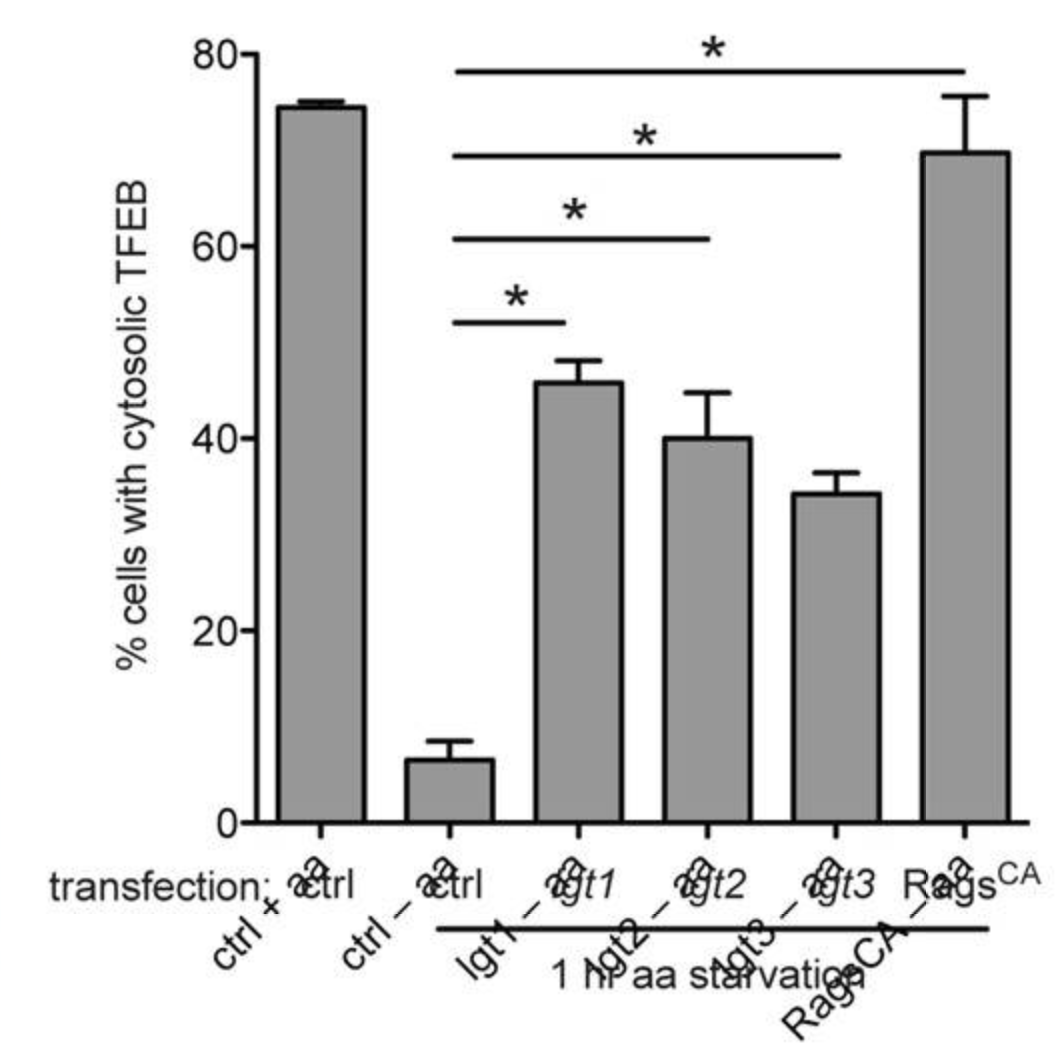
Figure 1. Amount of transcription factor EB (TFEB) found in the cell’s cytosol. Legionella uses Dot/Icm effectors to prevent TFEB from entering the nucleus, starting transcription, and using up amino acids needed by Legionella, as seen in the lgt1, lgt2, lgt3, and Rags genotypes.1
BSJ: Your paper mentions how the virulence of Legionella is dependent on a Dot/Icm type 4 secretion system and how this system is connected to the function of the mTORC1.1 What is the Dot/Icm type 4 secretion system and how is it related to mTORC1?
RV: The Dot/Icm secretion system is the primary system the bacterium uses to deliver bacterial proteins into host cells. You can think of it like a hypodermic needle. The bacterium proteins injected through this Dot/Icm secretion system end up manipulating the proteins of the mTORC1 master regulator (Fig 1.). Basically, Legionella is a devious organism. It’s delivering bacterial proteins to the inside of our cells, manipulating them into producing the amino acids Legionella needs to grow. If we make a mutant that lacks this secretion system, it cannot deliver any of these proteins into host cells. Consequently, the mutant ends up completely avirulent and cannot cause any disease.
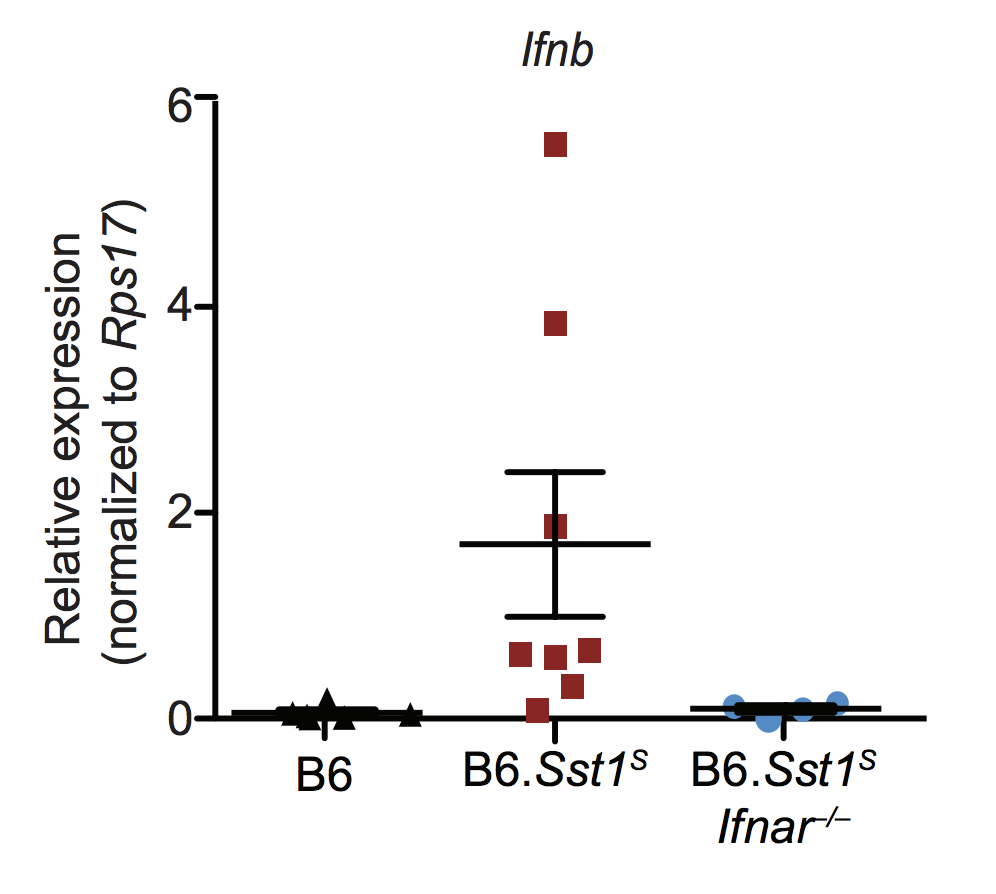
Figure 2. Interferon expression in wild type mice (B6), mice with a tuberculosis-susceptible mutation (B6.Sst1s), and mice with an additional mutation downregulating interferon response (B6.Sst1sIfnar-/-).2
BSJ: Interferons are signaling proteins released by a cell when a pathogen is present. In one of your papers you examined the elevated interferon protein response before tuberculosis progresses to an active disease.2 What were your results?
RV: For a long time we and many other labs have been interested in understanding the role of these type 1 interferons during tuberculosis. Type 1 interferons are generally considered to be very important when you’ve been infected with a virus. However, tuberculosis is caused by a bacterium. So it is very strange that the body would produce a lot of interferons because that doesn’t seem to be very helpful for a bacterial infection. In fact, there has been evidence showing that the production of these type of type1 interferons during tuberculosis is very detrimental to the host. The question we are trying to address is: why are the type1 interferons, that are normally helpful during a viral infection, detrimental during tuberculosis? That is a difficult problem to answer, since type 1 interferons regulate the expression of around a thousand different host genes. We focused on one particular protein that is induced by type 1 interferons, called the interleukin-1 receptor antagonist (IL1RA). What a student in my lab, Daisy Ji, found was that the production of this IL1 is very high in individuals affected by tuberculosis (Fig. 2). So, we think that part of the reason why type 1 interferons are bad for you when you’re infected by bacteria is that your body produces too much of IL1 receptor antagonist.
BSJ: You attempt to apply a lot of your work in Legionnaires’ disease to examining tuberculosis and how it functions. Have you found anything in your past research that you think will aid in your study of tuberculosis?
RV: The reason why we were interested in this protein IL1 receptor agonist is because we know that the function of that protein is to block this very important anti-bacterial protein called interleukin 1. We have been studying IL1 in Legionella for many years and we have shown that IL1 is crucial during Legionella infections. When we noticed that this antagonist of IL1 was produced during tuberculosis infections, it really caught our attention. I think it’s a great example of how something we learned while studying Legionella stimulated our thoughts when we studied tuberculosis. We realized that maybe tuberculosis was blocking this important IL1 response and that is one reason why those infections are more severe. So definitely, things that we are learning from one pathogen are helping us understand the other pathogen.
BSJ: You mentioned that vaccines for tuberculosis aren’t always efficacious. Given that tuberculosis is a bacterium, what makes it difficult to create a vaccine that is more efficient than the one we currently have?
RV: I would love to have an answer to that question, I think it’s probably the question in the tuberculosis field. We have developed effective vaccines for many infectious diseases, but for some it’s been extremely hard, and tuberculosis might be one of the hardest. We’ve made little progress creating an efficient vaccine for tuberculosis and fundamentally we don’t really understand why that is yet. I would argue that we need to do a lot more basic research into understanding how tuberculosis causes disease because we don’t yet fully understand how it avoids the immune response. That’s a very important question and we need more research to answer it.
References:
- De Leon JA, Qiu J, Nicolai CJ, Counihan JL, Barry KC, Xu L, Lawrence RE, Castellano BM, Zoncu R, Nomura DK, Luo ZQ, Vance RE (2017) Positive and Negative Regulation of the Master Metabolic Regulator mTORC1 by Two Families of Legionella pneumophila Effectors. Cell Reports, 21(8), 2031-2038. doi: 1016/j.celrep.2017.10.088.
- Ji DX, Chen KJ, Mukaida N, Kramnik I, Darwin KH, Vance RE (2018) Interleukin-1 receptor antagonist mediates type I interferon-driven susceptibility to Mycobacterium tuberculosis. Biorxiv, Aug 10 2018 (pre-print). doi: http://dx.doi.org/10.1101/389288.
- Price JV, Jiang K, Galantowicz A, Freifeld A, Vance RE (2018) Legionella pneumophila is directly sensitive to 2-deoxyglucose-phosphate via its UhpC transporter but is indifferent to shifts in host cell glycolytic metabolism. J Bacteriol, 200(16), e00176-18. doi:10.1128/JB.00176-18.
